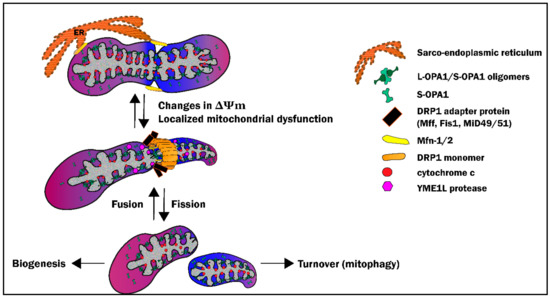
The crystal structure and biochemical characteristics of Mid51 suggested that it binds to ADP and GDP. Mff was discovered in siRNA screens, using cultured Drosophila DS2R+ cells, while Mid49/51 was found through analyses of mitochondrial proteomes. Steady state recruitment of Drp1 likely depends on other receptors such as Mff (mitochondrial fission factor) and Mid49/51 (MIEF1/2). However, Fis1 appears to recruit Drp1 in a subset of cell types, and/or under specific physiological conditions such as mitochondrial stress. A mammalian homolog of Fis1 has been identified. Analyses of dynamin homologs in Caenorhabditis elegans and in mammalian cells have identified Drp1 (a homolog of Dnm1), and shown that Dnm1/Drp1 are evolutionarily conserved division factors. Num1 and Mdm36 are unique, separate components with dual functions, connecting Dnm1 to the actin cortex and regulating both mitochondrial division and positioning within the cell. Yeast genetics was used to identify the central division component, Dnm1, and its receptor Fis1, and adaptors Mdv1, and Caf4. Mitochondrial receptors with transmembrane domains are involved in targeting Drp1 to the outer mitochondrial membrane. Loss of Drp1 results in long interconnected mitochondrial networks. It is a cytosolic dynamin-related GTPase, which moves to the mitochondrial outer membrane where it self assembles via GTP binding. Dnm1/Drp1 is the main component of mitochondrial division. Model organisms have been instrumental in identifying the core components of mitochondrial division and fusion. Identification of the molecular components has been determined in approximately the last 15 years ( Table 1). The dynamic nature of mitochondria was first observed during the early twentieth century. Defects in the core components of these systems, three dynamin-related GTPases, give rise to several disease conditions, including neonatal death with severe neural defects (defects in outer membrane protein Drp1, which mediates division), Charcot-Marie-Tooth neuropathy type 2A, a neurodegenerative disease of peripheral neurons, (defects in outer membrane protein Mfn2 which mediates fusion), and inherited forms of dominant optic atrophy (defects in inner membrane protein Opa1, which mediates fusion). These two opposing forces ensure that at any given time, the cell has a healthy mitochondrial population. In contrast, fusion is important for mixing of mitochondrial contents and maintaining electrical conductivity throughout the mitochondria. Division is crucial for maintaining the number of mitochondria in growing cells, regulating cell death pathways, and eliminating damaged mitochondria as part of quality control through mitophagy. In this review, we focus on mitochondrial division and fusion. There is dedicated protein machinery that controls the mitochondrial dynamics in the cell ( Table 1). Mitochondrial dynamics refer to the perpetual process of fusion, division, movement, and morphological changes which take place in response to the ever-changing physiological demands of cells. Mitochondrial fusion is regulated by proteosomal degradation of mitofusins, proteolytic processing of Opa1 and production of GTP. Outer membrane fusion is mediated by mitofusin while inner membrane fusion is mediated by Opa1. Mitochondrial fusion consists of outer membrane fusion and inner membrane fusion. Drp1 filaments constrict and divide mitochondria, working together with ER tubules and actomyosin filaments. On mitochondria, Drp1 assembles into helical filaments, wrapping around mitochondrial tubules. Mitochondrial division is initiated by recruitment of cytosolic Drp1 to the mitochondrial outer membrane by Drp1 receptors. Mitochondria continuously divide and fuse and control their morphology. Mitochondrial morphology is regulated by division and fusion
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
